Back MITF BS Microphthalmie-assoziierter Transkriptionsfaktor German Factor de transcripción asociado con microftalmia Spanish Microphthalmia-associated transcription factor French 小眼球症関連転写因子 Japanese MITF Ukrainian
Microphthalmia-associated transcription factor also known as class E basic helix-loop-helix protein 32 or bHLHe32 is a protein that in humans is encoded by the MITF gene.
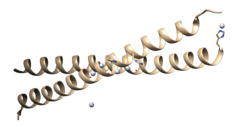
MITF is a basic helix-loop-helix leucine zipper transcription factor involved in lineage-specific pathway regulation of many types of cells including melanocytes, osteoclasts, and mast cells.[5] The term "lineage-specific", since it relates to MITF, means genes or traits that are only found in a certain cell type. Therefore, MITF may be involved in the rewiring of signaling cascades that are specifically required for the survival and physiological function of their normal cell precursors.[6]
MITF, together with transcription factor EB (TFEB), TFE3 and TFEC, belong to a subfamily of related bHLHZip proteins, termed the MiT-TFE family of transcription factors.[7][8] The factors are able to form stable DNA-binding homo- and heterodimers.[9] The gene that encodes for MITF resides at the mi locus in mice,[10] and its protumorogenic targets include factors involved in cell death, DNA replication, repair, mitosis, microRNA production, membrane trafficking, mitochondrial metabolism, and much more.[11] Mutation of this gene results in deafness, bone loss, small eyes, and poorly pigmented eyes and skin.[12] In human subjects, because it is known that MITF controls the expression of various genes that are essential for normal melanin synthesis in melanocytes, mutations of MITF can lead to diseases such as melanoma, Waardenburg syndrome, and Tietz syndrome.[13] Its function is conserved across vertebrates, including in fishes such as zebrafish[14] and Xiphophorus.[15]
An understanding of MITF is necessary to understand how certain lineage-specific cancers and other diseases progress. In addition, current and future research can lead to potential avenues to target this transcription factor mechanism for cancer prevention.[16]
- ^ a b c GRCh38: Ensembl release 89: ENSG00000187098 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000035158 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ Hershey CL, Fisher DE (April 2004). "Mitf and Tfe3: members of a b-HLH-ZIP transcription factor family essential for osteoclast development and function". Bone. 34 (4): 689–96. doi:10.1016/j.bone.2003.08.014. PMID 15050900.
- ^ Garraway LA, Sellers WR (August 2006). "Lineage dependency and lineage-survival oncogenes in human cancer". Nature Reviews. Cancer. 6 (8): 593–602. doi:10.1038/nrc1947. PMID 16862190. S2CID 20829389.
- ^ Hemesath TJ, Steingrímsson E, McGill G, Hansen MJ, Vaught J, Hodgkinson CA, et al. (November 1994). "microphthalmia, a critical factor in melanocyte development, defines a discrete transcription factor family". Genes & Development. 8 (22): 2770–80. doi:10.1101/gad.8.22.2770. PMID 7958932.
- ^ Ballesteros-Álvarez J, Dilshat R, Fock V, Möller K, Karl L, Larue L, et al. (3 September 2020). "MITF and TFEB cross-regulation in melanoma cells". PLOS ONE. 15 (9): e0238546. Bibcode:2020PLoSO..1538546B. doi:10.1371/journal.pone.0238546. PMC 7470386. PMID 32881934.
- ^ Pogenberg V, Ballesteros-Álvarez J, Schober R, Sigvaldadóttir I, Obarska-Kosinska A, Milewski M, et al. (January 2020). "Mechanism of conditional partner selectivity in MITF/TFE family transcription factors with a conserved coiled coil stammer motif". Nucleic Acids Research. 48 (2): 934–948. doi:10.1093/nar/gkz1104. PMC 6954422. PMID 31777941.
- ^ Hughes MJ, Lingrel JB, Krakowsky JM, Anderson KP (October 1993). "A helix-loop-helix transcription factor-like gene is located at the mi locus". The Journal of Biological Chemistry. 268 (28): 20687–90. doi:10.1016/S0021-9258(19)36830-9. PMID 8407885.
- ^ Cheli Y, Ohanna M, Ballotti R, Bertolotto C (February 2010). "Fifteen-year quest for microphthalmia-associated transcription factor target genes". Pigment Cell & Melanoma Research. 23 (1): 27–40. doi:10.1111/j.1755-148X.2009.00653.x. PMID 19995375. S2CID 43471663.
- ^ Moore KJ (November 1995). "Insight into the microphthalmia gene". Trends in Genetics. 11 (11): 442–8. doi:10.1016/s0168-9525(00)89143-x. PMID 8578601.
- ^ "MITF gene". Genetics Home Reference. National Institutes of Health, U.S. Department of Health & Human Services.
- ^ Lister JA, Robertson CP, Lepage T, Johnson SL, Raible DW (September 1999). "nacre encodes a zebrafish microphthalmia-related protein that regulates neural-crest-derived pigment cell fate". Development. 126 (17): 3757–67. doi:10.1242/dev.126.17.3757. PMID 10433906.
- ^ Delfgaauw J, Duschl J, Wellbrock C, Froschauer C, Schartl M, Altschmied J (November 2003). "MITF-M plays an essential role in transcriptional activation and signal transduction in Xiphophorus melanoma". Gene. 320: 117–26. doi:10.1016/s0378-1119(03)00817-5. PMID 14597395.
- ^ Ballotti R, Cheli Y, Bertolotto C (December 2020). "The complex relationship between MITF and the immune system: a Melanoma ImmunoTherapy (response) Factor?". Mol Cancer. 19 (1): 170. doi:10.1186/s12943-020-01290-7. PMC 7718690. PMID 33276788.