
Back تعرف جزيئي Arabic Molekularno prepoznavanje BS Molekülerkennung German Reconocimiento molecular Spanish Riconoscimento molecolare Italian 分子認識 Japanese 분자 인식 Korean Reconhecimento molecular Portuguese Молекулярное распознавание Russian Molekulsko prepoznavanje Serbo-Croatian
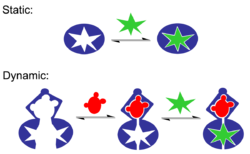
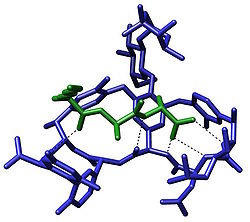


The term molecular recognition refers to the specific interaction between two or more molecules through noncovalent bonding such as hydrogen bonding, metal coordination, hydrophobic forces,[3][4] van der Waals forces, π-π interactions, halogen bonding, or resonant interaction[5] effects. In addition to these direct interactions, solvents can play a dominant indirect role in driving molecular recognition in solution.[6][7] The host and guest involved in molecular recognition exhibit molecular complementarity. Exceptions are molecular containers,[8][9] including, e.g., nanotubes, in which portals essentially control selectivity.[10][11][12][13] Selective partioning of molecules between two or more phases can also result in molecular recognition.[14] In partitioning-based molecular recognition the kinetics and equilibrium conditions are governed by the presence of solutes in the two phases.[15]
- ^ Knox JR, Pratt RF (July 1990). "Different modes of vancomycin and D-alanyl-D-alanine peptidase binding to cell wall peptide and a possible role for the vancomycin resistance protein" (Free full text). Antimicrobial Agents and Chemotherapy. 34 (7): 1342–1347. doi:10.1128/AAC.34.7.1342. PMC 175978. PMID 2386365.
- ^ Bielawski C, Chen Y, Zhang P, Prest P, Moore JS (1998). "A modular approach to constructing multi-site receptors for isophthalic acid". Chemical Communications (12): 1313–4. doi:10.1039/a707262g.
- ^ Lockett MR, Lange H, Breiten B, Heroux A, Sherman W, Rappoport D, et al. (July 2013). "The binding of benzoarylsulfonamide ligands to human carbonic anhydrase is insensitive to formal fluorination of the ligand". Angewandte Chemie. 52 (30): 7714–7717. doi:10.1002/anie.201301813. PMID 23788494. S2CID 1543705.
- ^ Breiten B, Lockett MR, Sherman W, Fujita S, Al-Sayah M, Lange H, et al. (October 2013). "Water networks contribute to enthalpy/entropy compensation in protein-ligand binding". Journal of the American Chemical Society. 135 (41): 15579–15584. CiteSeerX 10.1.1.646.8648. doi:10.1021/ja4075776. PMID 24044696. S2CID 17554787.
- ^ Cosic I (December 1994). "Macromolecular bioactivity: is it resonant interaction between macromolecules?--Theory and applications". IEEE Transactions on Bio-Medical Engineering. 41 (12): 1101–1114. doi:10.1109/10.335859. PMID 7851912. S2CID 23892544.
- ^ Baron R, Setny P, McCammon JA (September 2010). "Water in cavity-ligand recognition". Journal of the American Chemical Society. 132 (34): 12091–12097. doi:10.1021/ja1050082. PMC 2933114. PMID 20695475.
- ^ Baron R, McCammon JA (2013). "Molecular recognition and ligand association". Annual Review of Physical Chemistry. 64: 151–175. Bibcode:2013ARPC...64..151B. doi:10.1146/annurev-physchem-040412-110047. PMID 23473376.
- ^ Cram DJ, Cram JM (1997). Container molecules and their guests. Cambridge: Royal Society of Chemistry. ISBN 978-0-85186-972-8.
- ^ Brotin T, Dutasta JP (January 2009). "Cryptophanes and their complexes--present and future". Chemical Reviews. 109 (1): 88–130. doi:10.1021/cr0680437. PMID 19086781.
- ^ Lehn JM (1995). Supramolecular Chemistry. Weinheim: Wiley-VCH. ISBN 978-3-527-29312-4. OCLC 315928178.[page needed]
- ^ Gellman SH (August 1997). "Introduction: Molecular Recognition". Chemical Reviews. 97 (5): 1231–1232. doi:10.1021/cr970328j. PMID 11851448.
- ^ Chatterji D (2016). Basics of Molecular Recognition. Boca Raton, FL: CRC Press. ISBN 978-1-4822-1968-5.
- ^ Rotello V, Thayumanavan S, eds. (2008). Molecular recognition and polymers : control of polymer structure and self-assembly. Hoboken, N.J.: Wiley. ISBN 978-0-470-27738-6.
- ^ Bell TN, Feng K, Calvin G, Van Winkle DH, Lenhert S (October 2020). "Organic Composomes as Supramolecular Aptamers". ACS Omega. 5 (42): 27393–27400. doi:10.1021/acsomega.0c03799. PMC 7594120. PMID 33134702.
- ^ Zhou H, Shiel E, Bell T, Lin S, Lenhert S (November 2023). "Kinetic Mechanism of Surfactant-Based Molecular Recognition: Selective Permeability across an Oil–Water Interface Regulated by Supramolecular Aggregates". The Journal of Physical Chemistry B. 127 (47): 10201–10214. doi:10.1021/acs.jpcb.3c05017. PMID 37972386.