
Back تصميم الحمض النووي Arabic Dizajn nukleinskih kiselina Serbo-Croatian Dizajn nukleinskih kiselina Serbian
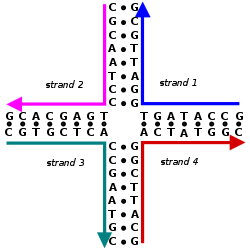
Nucleic acid design is the process of generating a set of nucleic acid base sequences that will associate into a desired conformation. Nucleic acid design is central to the fields of DNA nanotechnology and DNA computing.[2] It is necessary because there are many possible sequences of nucleic acid strands that will fold into a given secondary structure, but many of these sequences will have undesired additional interactions which must be avoided. In addition, there are many tertiary structure considerations which affect the choice of a secondary structure for a given design.[3][4]
Nucleic acid design has similar goals to protein design: in both, the sequence of monomers is rationally designed to favor the desired folded or associated structure and to disfavor alternate structures. However, nucleic acid design has the advantage of being a much computationally simpler problem, since the simplicity of Watson-Crick base pairing rules leads to simple heuristic methods which yield experimentally robust designs. Computational models for protein folding require tertiary structure information whereas nucleic acid design can operate largely on the level of secondary structure. However, nucleic acid structures are less versatile than proteins in their functionality.[2][5]
Nucleic acid design can be considered the inverse of nucleic acid structure prediction. In structure prediction, the structure is determined from a known sequence, while in nucleic acid design, a sequence is generated which will form a desired structure.[2]
- ^ Mao, Chengde (December 2004). "The Emergence of Complexity: Lessons from DNA". PLOS Biology. 2 (12): 2036–2038. doi:10.1371/journal.pbio.0020431. ISSN 1544-9173. PMC 535573. PMID 15597116.
- ^ a b c Cite error: The named reference
Dirks04was invoked but never defined (see the help page). - ^ Cite error: The named reference
Seeman82was invoked but never defined (see the help page). - ^ Cite error: The named reference
Sherman06was invoked but never defined (see the help page). - ^ Cite error: The named reference
Brenneman02was invoked but never defined (see the help page).